PCACORR: A Simple Method to Improve RXTE PCA Spectra
A simple method has been developed to improve the quality of RXTE PCA spectra by appliying an empirical correction based on the analysis of the Crab spectra. By fitting at once several hundred of PCA spectra of the Crab to a simple power-law model a composite spectrum with ~109 counts can be achieved. The combined ratio spectrum reveals many instrumental features not included in the current calibration. These are then used as scaling factors to correct any PCA spectra reducing the contribution of these residuals and significantly improving the fits.
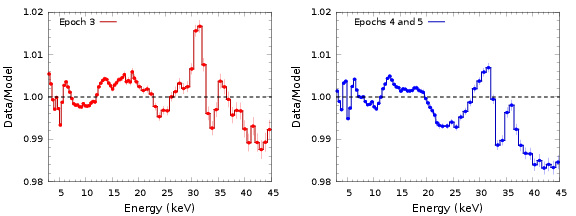
Correction curves are derived for each one of the 5 PCUs. Becuase of the important changes in gain between epochs, a distinction is made in the correction for data taken during epoch 3, and those taken during epochs 4 and 5. Note that epochs 1 and 2 are not covered at all. Also, different corrections are provided for observations that are reduced by including all layers and observations that only take data from the first layer.
We provide a script that can be used to apply this correction to any PCA spectra, along with the corresponding correction curves.
| File | Description | Version | Date |
| pcacorr.py | Correction script | 0.1d | Sep 2, 2014 |
| corrections.tgz | Correction files | 0.1d | Nov 23, 2018 |
The PCACORR script is written in Python. It requires the numpy and pyfits packages. Both can be easily install using the PIP installer (e.g. pip install pyfits).
The PCACORR Script
| Usage: | pcacorr.py [options] PREFIX | |
|
Produce a new observation file using the correction factors derived from
the analysis to the Crab (see Garcia et al Apj...). The response corresponding to the observation is require. The response file name is obtained from the RESPFILE key in the observation header or it must be supplied. The correction file is also required.
PREFIX can be a single observation file or a group (e.g. *.pha). Please check the
gain epoch of your observation!
*** Epoch 3 (MJD): 50188-51259 *** Epochs 4 & 5 (MJD): > 51259 |
||
| Options: | ||
| -h, | --help | show this help message and exit |
| -v, | --version | show version number |
| -r RESPFILE, | --response=RESPFILE | specify response file |
| -p PCU, | --pcu=PCU | specify which PCU (0-4, default 2) |
| -l LAYER, | --layer=LAYER | specify if observation comes from the first layer ("1") or all ("a", default) |
| -e EPOCH, | --epoch=EPOCH | specify if observation was taken during gain epoch 3 only ("3o"), or during epochs 4 or 5 ("45", default) |
| -c CORFILE, | --correction=CORFILE | specify correction file (default corr_pcu2_e45_la.out). Notice that this option overrides -p, -l, and -e options |
| -o OUTFILE, | --output=OUTFILE | specify alternative output file |
| -b, | --background | Use associated background file. In this case the corrected file will contain (C-B)/x + B, where C are the cts of the original spectrum, B are the cts of the background, and x is the correction factor. Be aware that the BACKFILE key must be defined in the spectrum file with the correct background filename |
Examples:
Correcting observation 'spectrum.pha', taken with PCU-2, during gain Epoch 4, with all layers (just use the defaults!):
./pcacorr.py spectrum.pha
Correcting observation 'spectrum.pha', taken with PCU-0, during gain Epoch 3, with only the first layer:
./pcacorr.py -p 0 -e 3o -l 1 spectrum.pha
Do the same by specifiying the desired correction file:
./pcacorr.py -c corr_pcu0_e3o_l1.out spectrum.pha
If you want the corrected observation file to have a particular name (only works for single files):
./pcacorr.py -o super-nice-spectrum.pha spectrum.pha
In the case that your observation in not background corrected, use the -b option to tell the script to look for the corresponding background file:
./pcacorr.py -b spectrum.pha
Finally, you can use all these options (except -o) for all the observations in your folder:
./pcacorr.py -b -pcu 4 -e 45 -l a *.pha
*** Important ***: please note that this script will apply the correction curve according to the options supplied by the user, but it does not checks if they are appropriate for the given observation. So be careful in double checking your data's gain epoch, PCU, and layer options. This is particularly important when running the script on a group of files.
Individual correction files: All Layers
Individual correction files: Layer 1
This is the first release of this method, so having feedback from potential users is important. Both the PCACORR script and the correction files are likely to change in the future. If you wish to be notified of such changes,or have any comments or questions, please feel free to contact me!